Strand on a real TP53 map
This page uses a real TP53 deep mutational scanning scoreset (urn:mavedb:00001213-a-1) from MaveDB to show, concretely, how Strand behaves when you ask it to design a focused, mechanism-rich library rather than a brute-force screen.
1. From a dense TP53 map to a library design
We start from a dense functional map: a TP53 deep mutational scan where each missense variant has an experimental score. Strand ingests this scoreset via the MaveDB Python ecosystem, normalizes it into a canonical perturbation library, and annotates each variant with:
- A coarse functional effect class (severe loss-of-function, partial loss-of-function / hypomorph, WT-like, assay-specific gain-of-function)
- Domain labels across the TP53 structure (for this scoreset, primarily DNA-binding core and adjacent linker; the same scheme supports full-length scans)
- Coarse position windows (e.g.,
pos_100_149) for region-aware coverage
The same pattern extends to other perturbation types (MPRA, base-editor tiling, CRISPR tiling) as long as we can map them into a shared perturbation schema.
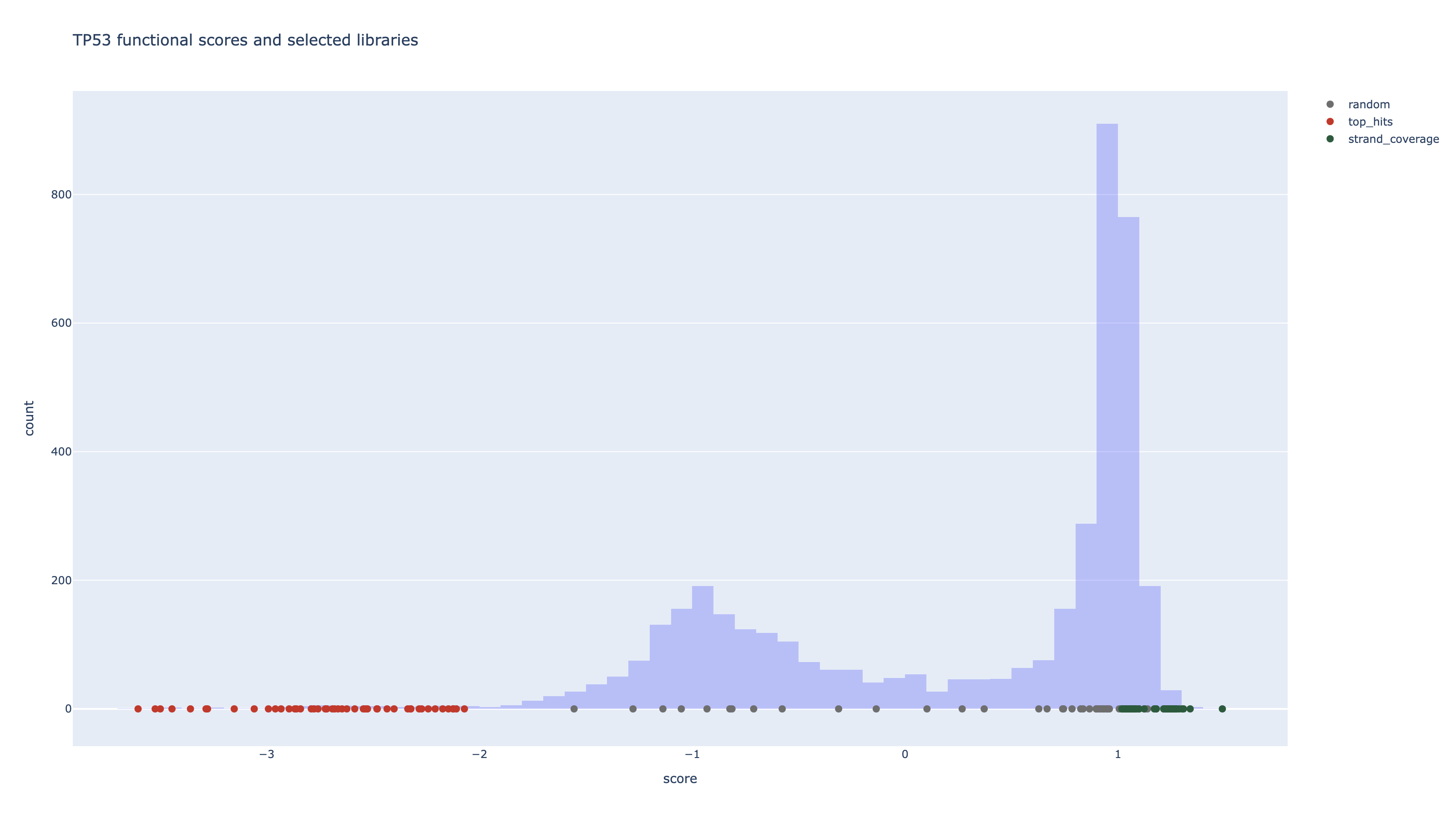
2. Three library designs on the same TP53 map
To illustrate how Strand behaves on a real dataset, we compare three strategies for selecting a library of K = 48 perturbations:
- Random: sample variants uniformly from the scoreset.
- Top-hits: pick the most extreme loss-of-function variants by score.
- Strand coverage-aware: optimize for high functional impact while explicitly covering different mechanisms and domains.
Mechanism and Domain Coverage
| Strategy | K | Mechanism Coverage (bins → count) | Domain Coverage (domains → count) | Assay Mix (modules → count) | Rough Assay Cost (units) | Score Range (min / median / max) |
|---|---|---|---|---|---|---|
| random | 48 | wt_like: 25, gain_of_function: 12, moderate_lof: 7, severe_lof: 4 | DBD_core: 46, linker: 2 | DSF_SEC: 39, FUNCTIONAL: 9 | 105.0 (mean 2.19) | -1.555 / 0.900 / 1.139 |
| top hits | 48 | severe_lof: 48 | DBD_core: 46, linker: 2 | FUNCTIONAL: 46, DSF_SEC: 2 | 142.0 (mean 2.96) | -3.604 / -2.672 / -2.070 |
| strand coverage | 48 | gain_of_function: 41, wt_like: 7 | DBD_core: 25, linker: 23 | DSF_SEC: 48 | 96.0 (mean 2.00) | 1.019 / 1.217 / 1.490 |
The Strand coverage-aware panel preserves strong hits but deliberately spends slots on under-represented mechanisms and domains so that each experiment is maximally informative for downstream modeling.
3. What actually ends up in the Strand panel
Below are a few example entries from the Strand coverage-aware panel. Each row is a concrete perturbation Strand would propose for a library, with its HGVS label, domain, and mechanism bin.
| HGVS Protein | Pos | AA Change | Score | Mechanism Bin | Domain | Region Bin |
|---|---|---|---|---|---|---|
| NP_000537.3:p.Leu130Tyr | 130 | Leu→Tyr | 1.490 | gain_of_function | DBD_core | pos_100_149 |
| NP_000537.3:p.Leu299Ter | 299 | Leu→Ter | 1.038 | wt_like | linker | pos_250_299 |
| NP_000537.3:p.Gly266Phe | 266 | Gly→Phe | 1.339 | gain_of_function | DBD_core | pos_250_299 |
| NP_000537.3:p.Glu298Ter | 298 | Glu→Ter | 1.246 | gain_of_function | linker | pos_250_299 |
| NP_000537.3:p.Lys164Asp | 164 | Lys→Asp | 1.308 | gain_of_function | DBD_core | pos_150_199 |
| NP_000537.3:p.Leu145Ala | 145 | Leu→Ala | 1.301 | gain_of_function | DBD_core | pos_100_149 |
| NP_000537.3:p.His296Ter | 296 | His→Ter | 1.180 | gain_of_function | linker | pos_250_299 |
| NP_000537.3:p.Arg174Ter | 174 | Arg→Ter | 1.287 | gain_of_function | DBD_core | pos_150_199 |
| NP_000537.3:p.Glu294Ter | 294 | Glu→Ter | 1.178 | gain_of_function | linker | pos_250_299 |
| NP_000537.3:p.Pro278Ile | 278 | Pro→Ile | 1.277 | gain_of_function | DBD_core | pos_250_299 |
| NP_000537.3:p.Glu294Ter | 294 | Glu→Ter | 1.171 | gain_of_function | linker | pos_250_299 |
| NP_000537.3:p.Arg174Ter | 174 | Arg→Ter | 1.271 | gain_of_function | DBD_core | pos_150_199 |
| NP_000537.3:p.Thr304Ter | 304 | Thr→Ter | 1.123 | gain_of_function | linker | pos_300_349 |
| NP_000537.3:p.Met133Phe | 133 | Met→Phe | 1.269 | gain_of_function | DBD_core | pos_100_149 |
| NP_000537.3:p.His297Ter | 297 | His→Ter | 1.098 | gain_of_function | linker | pos_250_299 |
| NP_000537.3:p.Gly279Tyr | 279 | Gly→Tyr | 1.268 | gain_of_function | DBD_core | pos_250_299 |
4. How this generalizes beyond TP53
This TP53 slice is a concrete example of one product surface for Strand: functional genomics on a small number of known targets. In a typical engagement, a team brings:
- 3–5 genes / pathways they care about
- Existing functional data (DMS, MPRA, CRISPR tiling) or model predictions
- Assay and cost constraints across multiple readouts and cellular contexts
Strand ingests those functional maps, annotates perturbations with domain, mechanism, and region labels, and designs focused libraries that:
- Span loss-of-function, hypomorph, gain-of-function, splice and regulatory mechanisms
- Cover key exons, domains, and regulatory regions instead of over-sampling a single hotspot
- Respect constraints on assay complexity, number of cells, and readout throughput
The same optimization loop can then be applied to:
- CRISPR tiling guides across a regulatory region
- Pairwise perturbations for synergy and resistance mapping
- Context-specific libraries across different cell types or treatments
Every experiment returns labeled data that feeds back into the scoring and selection models, so the next library is strictly smarter. This page is one vertical slice showing what that looks like for TP53; the same pattern is designed to generalize across a portfolio of targets where you care about making each experiment slot carry as much information as possible.